Background
A successful CITRUS run will produce results that indicate that features of certain clusters of cell populations within the data are significantly different between groups of samples and may be important to understanding the underlying differences in these sample groups (e.g. responders versus non-responders). The CITRUS results package provides basic information to understand the cell phenotype that corresponds to the significant clusters and other data about these clusters, but most investigators will want greater freedom to explore and analyze the events that CITRUS has isolated in other ways. In order to accommodate this, the Cytobank platform supports the export of CITRUS clusters as new FCS files for any desired downstream analysis
The events within any CITRUS cluster can be exported into a new FCS file.
Use the links below to jump to any section in the article:
- How to Export Clusters from CITRUS
- Exported Clusters Experiment Naming and Navigation
- Exported Cluster FCS Files - Anatomy
- Limitations of Export Clusters with Compensation
- Illustrations generated after CITRUS cluster export
How to Export Clusters from CITRUS
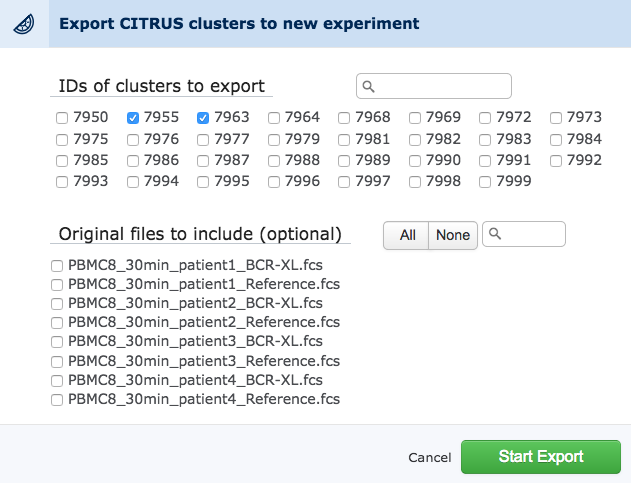
After a CITRUS run has completed, a button to export clusters will appear on the CITRUS setup page. Clicking this button will make a popup dialog appear that lists the clusters from the CITRUS run by ID. Files from the original experiment will also be included. Use this dialog to select desired clusters for export and then click to start the export. The export is a background task and the page can be closed or otherwise navigated away from and returned to later. Remember that the same events will appear in multiple clusters, so it makes sense to be tactical about which clusters to export so as not to create confusion in downstream analysis.
When a CITRUS run completes, a button will appear on the setup page to export clusters.
Once clicked, the export clusters button will make a dialog appear where clusters and original files can be chosen.
Exported Clusters Experiment Naming and Navigation
When the cluster export is finished, a new experiment will be created to house the resulting FCS files. This experiment will appear in the Experiment Manager and have a name that indicates it is based on exported clusters, along with which CITRUS run it originated from. For example, a CITRUS run named "Demo CITRUS run" will be named "CITRUS - Demo CITRUS run - Exported Clusters - 1" when it is exported. The number at the end will change depending on how many experiments have been exported from the same CITRUS run, since there can be many.
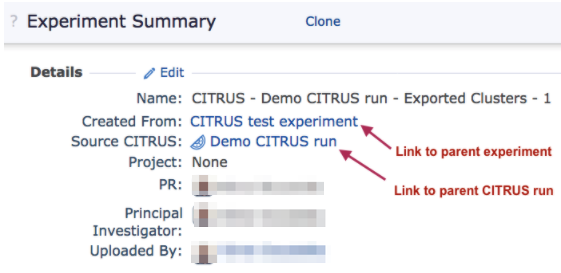
The exported experiment will be automatically linked to its parent experiment or parent CITRUS run and can be easily found on the linked experiments tree on the experiment summary page:
The exported clusters experiment links back to its parent CITRUS run and parent experiment.
The exported clusters experiment will also be tracked within the parent CITRUS run. A list of exported clusters experiments is displayed below the export clusters button:
A list of experiments created from exporting clusters is maintained within the CITRUS run.
Exported Cluster FCS Files - Anatomy
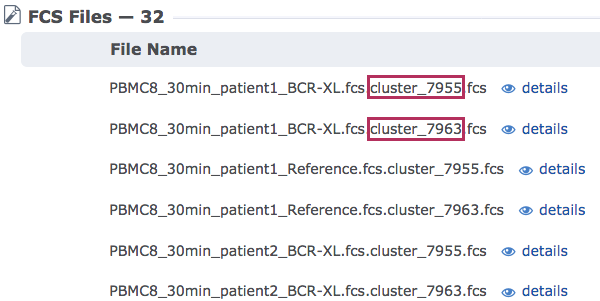
The new FCS files in the experiment will indicate which cluster they originated from within their file names. This can be seen anywhere the filename is displayed. An overview of the files in the experiment can be viewed on the experiment summary page under the FCS files list:
The list of files on the experiment summary page of the exported clusters experiment - filenames indicate originating cluster.
Limitations of Export Clusters with Compensation
When raw data are passed into CITRUS for analysis, they are scaled and compensated according to the settings at the time of the run. When clusters are exported this process has to be reversed to return the data to their raw state from the scaled and compensated version. As it stands currently, the scaling applied to the data originally can be reversed but the compensation cannot. Thus, if compensation was used for the CITRUS run, the FCS files exported from clusters in this run will appear compensated even when no compensation matrix is being applied to them. Put another way, the data are permanently compensated. Please keep this in mind when making comparisons of the exported clusters and the raw data, so as not to doubly compensate the exported clusters. Note: contact Cytobank Support if this is problematic for your workflow.
Illustrations generated after CITRUS cluster export
After exporting CITRUS clusters, two illustrations will be generated automatically: a heatmap and box plots. You can access these illustrations from the Illustration list on the Experiment Summary page.
CITRUS clusters and CITRUS groups will be automatically generated as Figure dimensions that could be used to create other meaningful Illustrations.
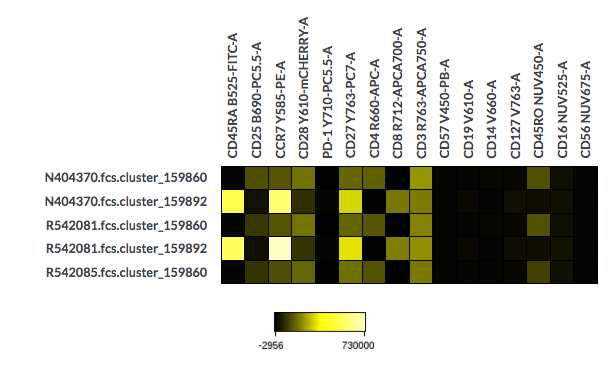
CITRUS heatmap
The raw median of the CITRUS clustering channels will be displayed for the first 5 new FCS files generated.
Example of the saved CITRUS heatmap in the created experiment after the cluster export.
See more on How to create and modify a Heatmap.
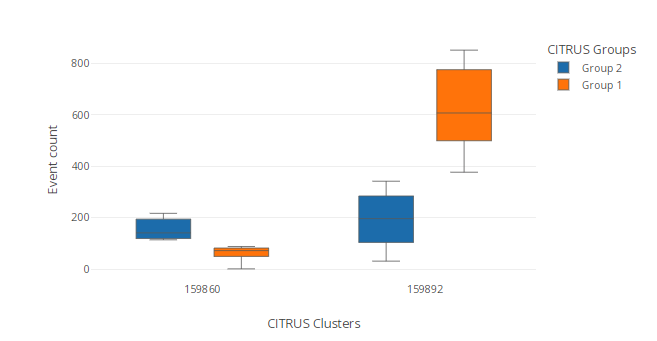
Box plot
CITRUS clusters will be grouped in box plots as X-axis groups, and CITRUS groups (the ones that were compared on the CITRUS run) will be assigned as Subgroups. See more on Summary plots generation and Statistical inference methods to modify the default Box plot.
Example of the saved CITRUS Box plot in the created experiment after the cluster export.
Tip for Success
If you want to export the original files together with the CITRUS clusters of interest, but the files have an unequal number of events, export the central CITRUS cluster instead when equal event sampling was selected on the CITRUS run.