Background
Dimensionality reduction algorithms tSNE-CUDA, opt-SNE, and UMAP offer the opportunity to perform analysis far faster than viSNE. Many analyses will complete in seconds to minutes, making run time negligible in your overall analysis workflow. To learn about how fast or slow your analysis may run, we:
- Compare the run time for the different algorithms in the DRS
- Show how some of the tunable parameters impacted run time in our analysis. See this article to learn about the impact of algorithm parameters on viSNE run times.
Run times of tSNE-CUDA, opt-SNE and UMAP
Using the default settings in the Cytobank platform for each algorithm, here is a comparison of the run time of the three algorithms on the same data set. Results may vary across data sets.
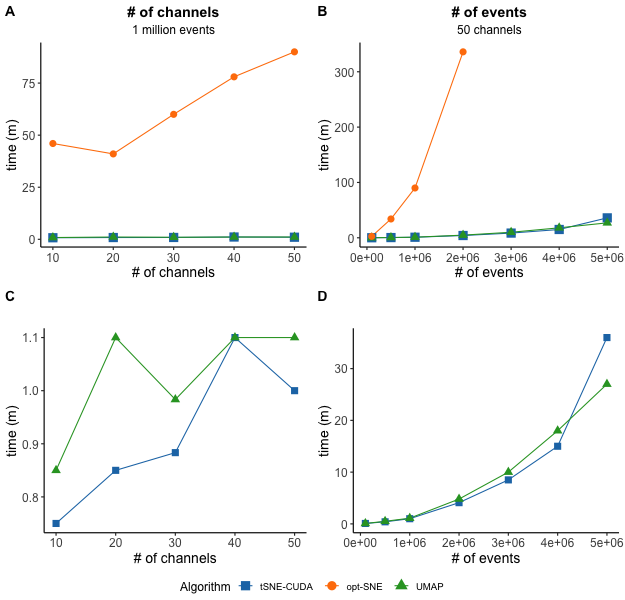
(Impact of data dimensions on algorithm run time in the DRS. The size of the data (events and channels) is generally the most important factor in determining algorithm run time, as compared to other algorithm settings. Run time is shown for 10, 20, 30, 40, or 50 channels using 1 million events for all three algorithms (A) and only tSNE-CUDA and UMAP (C). The right plots show run time for 50 channels of 100,000, 500,000, 1M, 2M, 3M, 4M and 5M events for all three algorithms (B) and only tSNE-CUDA and UMAP (D). Note, at this time, for a 50-channel file, opt-SNE can only accommodate only up to 2 million events, while tSNE-CUDA and UMAP are capped at 10 million events in the Cytobank platform. tSNE-CUDA and UMAP completed 1 million event data sets with up to 50 channels in around 1 minute, while opt-SNE took between 46 to 90 minutes. To assess both a high number of events (5 million) and channels (50), tSNE-CUDA took 36 minutes and UMAP took 27 minutes. The largest opt-SNE tested here was 2 million events and 50 channels and completed in 5.6 hours.)
Default settings used
Impact of dimensionality reduction settings on run time
tSNE-CUDA
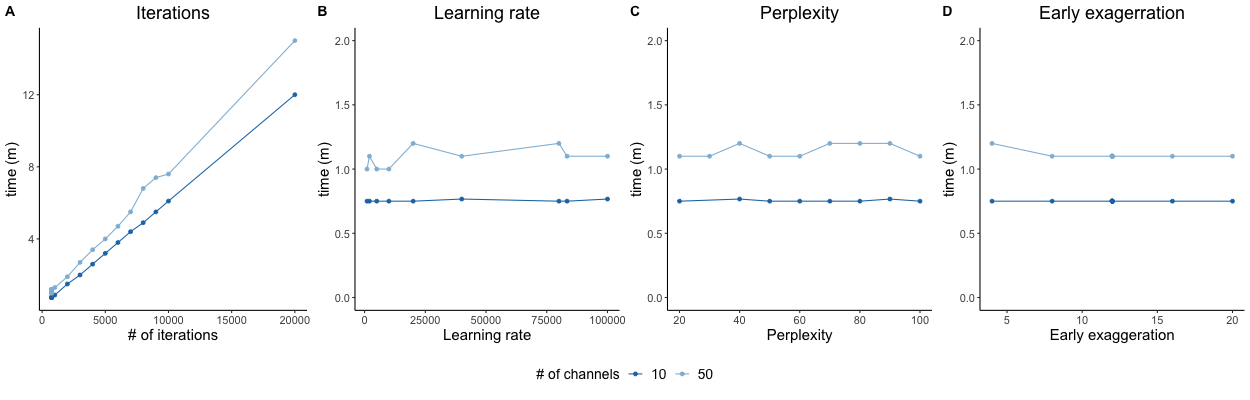
In tSNE-CUDA, the user specifies how many iterations, or repetitions, are performed. As one might expect, the number of iterations is correlated directly with run time. The other parameters (learning rate, perplexity, and early exaggeration) have minimal impact on run time in our test data.
(tSNE-CUDA settings and run times. Run time is shown for a 1 million event data set of either 10 or 50 channels, when adjusting iterations (A), learning rate (B), perplexity (C), or early exaggeration (D) parameters and keeping others as the default.)
opt-SNE
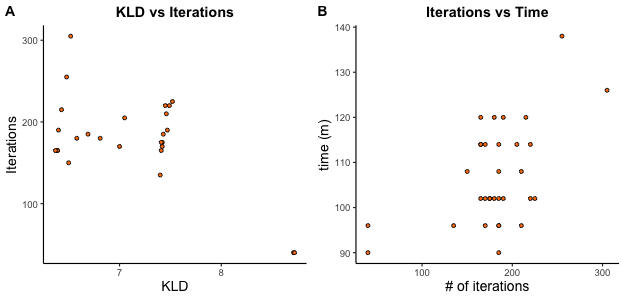
In contrast to tSNE-CUDA, opt-SNE tunes itself over the course of embedding, adjusting the length of the early exaggeration phase, and the total number of iterations. opt-SNE stops running when the embedding is optimal, as determined by a plateauing KLD. Since iteration number cannot be controlled for, it is challenging to compare other parameters to run time. We observe that for the same data set, comprising 1 million events and 50 channels, running with different combinations of tunable parameters early exaggeration, perplexity, and learning rate, the KLD tended to inversely correlate with the number of iterations (A). In other words, a high final KLD error rate, a measure of a lower quality result, was associated with stopping the run earlier, after fewer iterations. Lower KLD values, a surrogate for a better quality result, were associated with a higher number of iterations. As expected, a higher number of iterations is generally associated with a longer run time (B).
(opt-SNE settings and run times for a data set of 1 million events and 50 channels)
UMAP
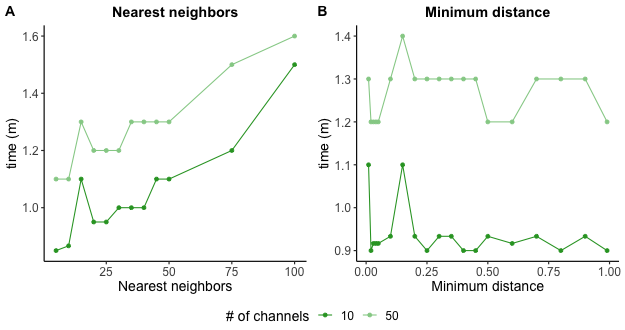
UMAP run times tended to increase with higher nearest neighbor values and was not impacted by changing minimum distance.
(UMAP settings and run times)