Background
When using the Cytobank API, there are many times that specific IDs are required (i.e. experiment ID, FCS file ID, SPADE ID, viSNE ID, etc.). These IDs can also be useful even when using the Cytobank GUI. This guide will take you through understanding URLs in Cytobank in order to gather ID information for any request that may require it.
Basic Concepts
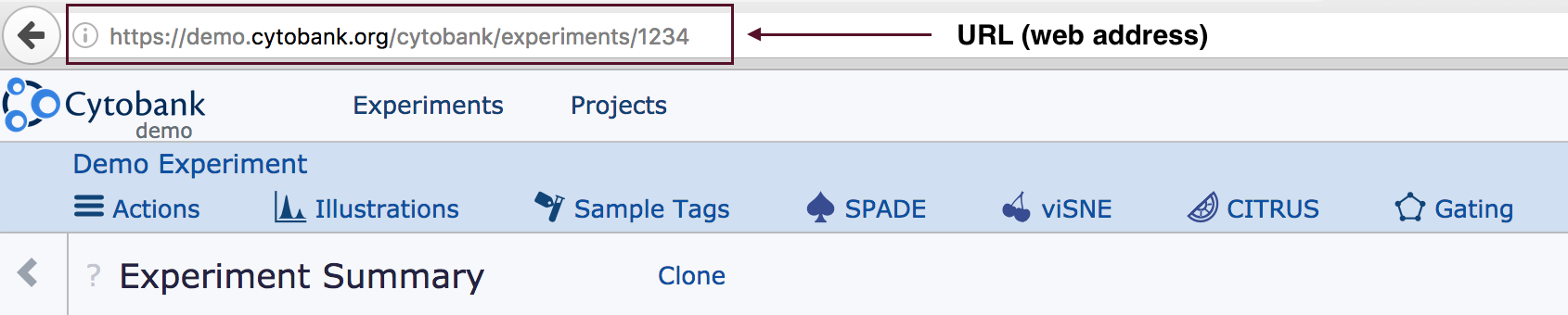
- Locating the URL (also known as the web address)
- This is typically found at the top of the web browser.
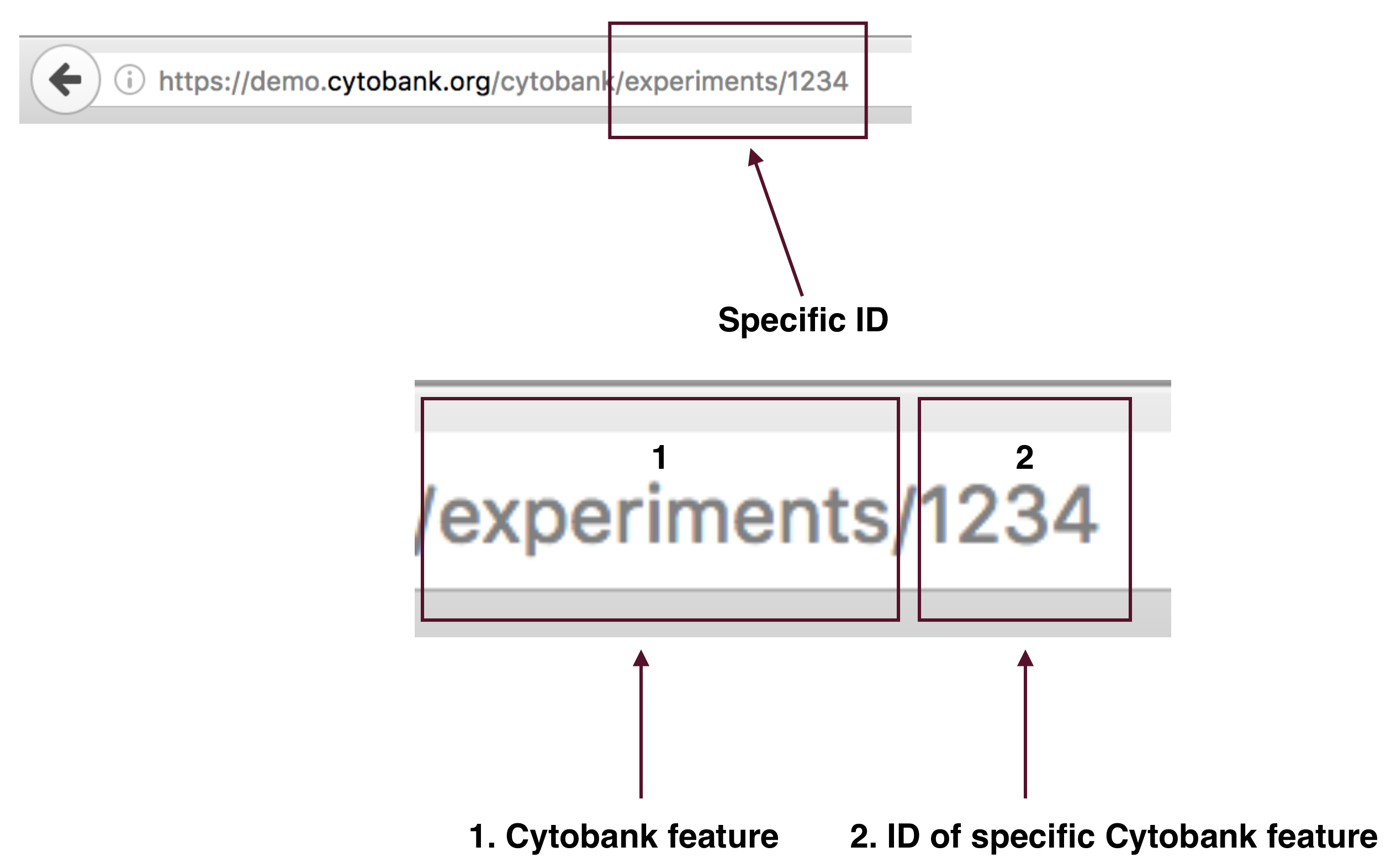
- Locating specific IDs
- There are two parts to a specific ID:
- The Cytobank feature
- The ID of the specific Cytobank feature
A More Complex Example
In the above Basic Concepts URL, we are examining the Experiments feature, that contains the ID 1234. We associate the specific Cytobank feature with a unique feature ID. URLs can contain many IDs that pertain the different Cytobank features.
For example, we may have created a viSNE analysis within Cytobank and are provided with the following URL:
https://demo.cytobank.org/cytobank/experiments/1234/visne_advanced_analyses/5678
We see that there are now 2 Cytobank features, an experiment ID [blue], as well as a viSNE ID [red]. It is important to remember that each ID is unique to its own Cytobank feature.
In Practice
Within the CytobankAPI R package, we see various endpoints that require IDs to be passed with each endpoint call. We will now walk through an example of using Cytobank feature IDs in combination with the CytobankAPI R package based on the examples above.
Recall that from the above examples, we currently have the following feature IDs:
- Experiment ID: 1234
- viSNE ID: 5678
If we would like to look up details about a specific experiment, we can utilize the CytobankAPI::experiments.show() function. This function takes in two parameters, one Cytobank API token (learn how to generate a Cytobank API token), and one ID parameter, experiment_id. We can utilize our previously gathered experiment ID for the experiment_id parameter.
CytobankAPI::experiments.show(UserSession=cyto_session, experiment_id=1234)
We have now seen how to utilize these IDs within the CytobankAPI R package.